Contributed by Poyin Chen
In the spirit of Halloween, the blog post today will be on the genetic difference of predatory bacteria.
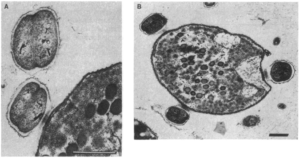
Vampirococcus is a predatory bacteria found in lakes in Spain and feeds on Chromatium, a purple sulfur bacteria. As the name suggests, Vampirococcus gets its food by sucking out the insides of its prey (Figure 1). Unfortunately, Vampirococcus has never before been grown in culture. As a consequence, we have no information on the genetic makeup of this organism. (1)
Many other predatory bacteria do exist and have been isolated in pure culture, allowing for genome sequencing and comparative analysis against non-predatory bacteria. When compared against non-predators, predatory bacteria were found to encode more genes for mevalonate metabolism, adhesion and signaling, and essential metabolite scavenging. In contrast, non-predators had more genes dedicated toward DOXP metabolism, amino acid biosynthesis, and riboflavin biosynthesis. Additionally, predatory bacteria were found to encode significantly more proteases, the majority of which are involved in protein and membrane degradation (2).
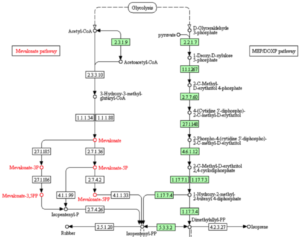
The pronounced differences in gene function abundances between predatory and non-predatory bacteria indicate that predatory bacteria are able to co-opt metabolites found in the prey cytosol and incorporate these metabolites into their own, incomplete metabolic processes. This is observed in the predatory preference for mevalonate metabolism from Glycolysis while non-predators favor the alternate pathway (DOXP) stemming from Glycolysis (Figure 2) (2).
Knowing what we know about bacterial vampires, one begins to wonder if, like their bacterial counterparts, human vampires need to drink our blood to harvest components such as white blood cells to keep themselves disease free. The more you know…
Figure 1
Image source: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC323246/pdf/pnas00311-0181.pdf
Figure 2
Image source: http://www.genome.jp/kegg/pathway.html
References:
- Guerrero R, Pedros-Alio C, Esteve I, Mas J, Chase D, Margulis L. 1986. Predatory prokaryotes: predation and primary consumption evolved in bacteria. Proc Natl Acad Sci U S A 83:2138-2142.
- Pasternak Z, Pietrokovski S, Rotem O, Gophna U, Lurie-Weinberger MN, Jurkevitch E. 2013. By their genes ye shall know them: genomic signatures of predatory bacteria. ISME J 7:756-769.